Mean Sea Surface Temperature#
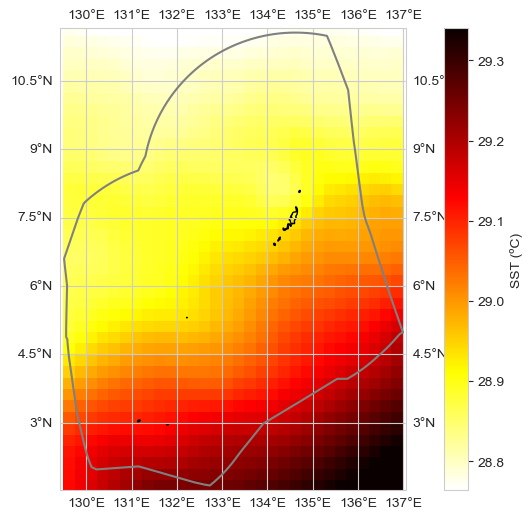
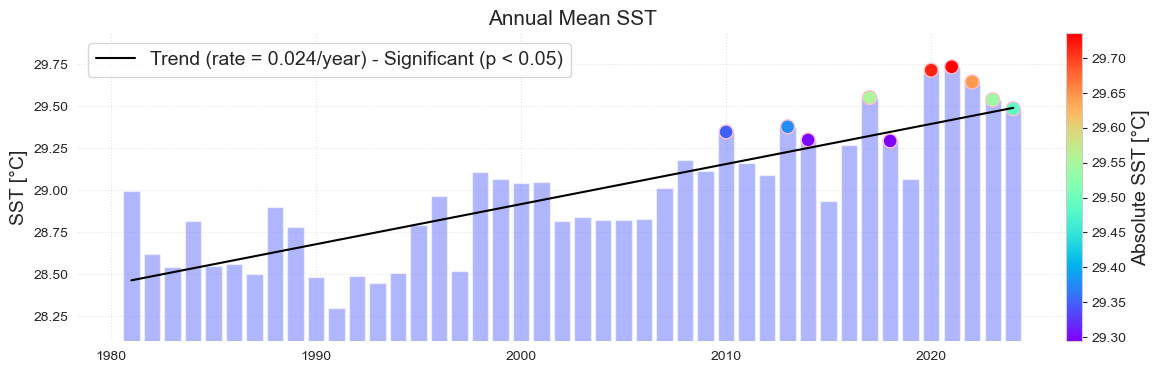
Figure. Sea Surface Temperature (SST) Change from satellite. The map (top) shows the change in mean SST (°C) in the vicinity of Palau over the period 1982–2024 from the NOAA OISSTv2 satellite. The grey line is the Palau EEZ. The bar plot (bottom) shows the mean SST averaged over the area within the top plot. The trend is statistically significant (p < 0.05). The colored dots represent the 10 warmest years on record.
Setup#
First, we need to import all the necessary libraries. Some of them are specifically developed to handle the download and plotting of the data and are hosted at the indicators set-up repository in GitHub
https://www.ncei.noaa.gov/products/optimum-interpolation-sst
Show code cell source
import warnings
warnings.filterwarnings("ignore")
import sys
import os
import os.path as op
import xarray as xr
import geopandas as gpd
import pandas as pd
import cartopy.crs as ccrs
import numpy as np
import matplotlib.pyplot as plt
from myst_nb import glue
sys.path.append("../../../../indicators_setup")
from ind_setup.plotting import plot_base_map, plot_bar_probs, plot_map_subplots, fontsize
from ind_setup.plotting_int import plot_timeseries_interactive, plot_oni_index_th
sys.path.append("../../../functions")
from data_downloaders import download_oni_index
from ocean import process_trend_with_nan
lon_site, lat_site = 134.368203,7.322074
#Area of interest
lon_range = [129.4088, 137.0541]
lat_range = [1.5214, 11.6587]
shp_f = op.join(os.getcwd(), '..', '..','..', 'data/Palau_EEZ/pw_eez_pol_april2022.shp')
shp_eez = gpd.read_file(shp_f)
path_data = "../../../data"
path_figs = "../../../matrix_cc/figures"
data = data_xr = xr.load_dataset(op.join(path_data, 'sst_daily_1981_2024_palau.nc'))
dataset_id = 'sst'
Analysis#
fig, ax = plot_base_map(shp_eez = shp_eez, figsize = [10, 6])
im = ax.pcolor(data.lon, data.lat, data.mean(dim='time')[dataset_id], transform=ccrs.PlateCarree(),
cmap = 'hot_r', vmin = np.percentile(data.mean(dim = 'time')[dataset_id], 1),
vmax = np.percentile(data.mean(dim = 'time')[dataset_id], 99))
ax.set_extent([lon_range[0], lon_range[1], lat_range[0], lat_range[1]], crs=ccrs.PlateCarree())
plt.colorbar(im, ax=ax, label='SST (ºC)')
glue("average_map", fig, display=False)
plt.savefig(op.join(path_figs, 'F12_SST_map.png'), dpi=300, bbox_inches='tight')

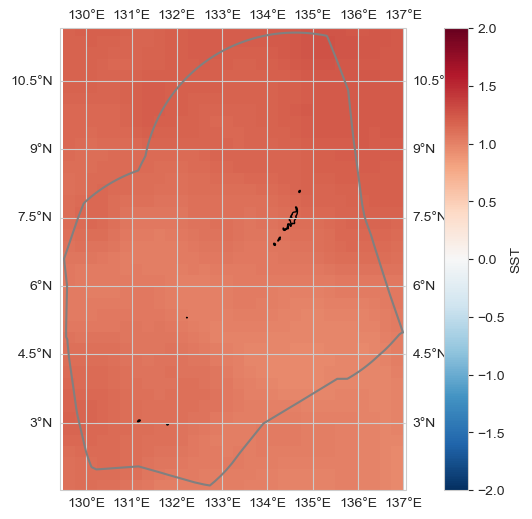
Change#
trend_m, _, _, _, _ = process_trend_with_nan(data[dataset_id])
fig, ax = plot_base_map(shp_eez = shp_eez, figsize = [10, 6])
im = ax.pcolor(data_xr.lon, data_xr.lat,
trend_m,
transform=ccrs.PlateCarree(),
cmap = 'RdBu_r',
vmin = -2,
vmax = 2,
)
ax.set_extent([lon_range[0], lon_range[1], lat_range[0], lat_range[1]], crs=ccrs.PlateCarree())
plt.colorbar(im, ax=ax, label= 'SST')
plt.savefig(op.join(path_figs, 'F12_SST_map_trend.png'), dpi=300, bbox_inches='tight')
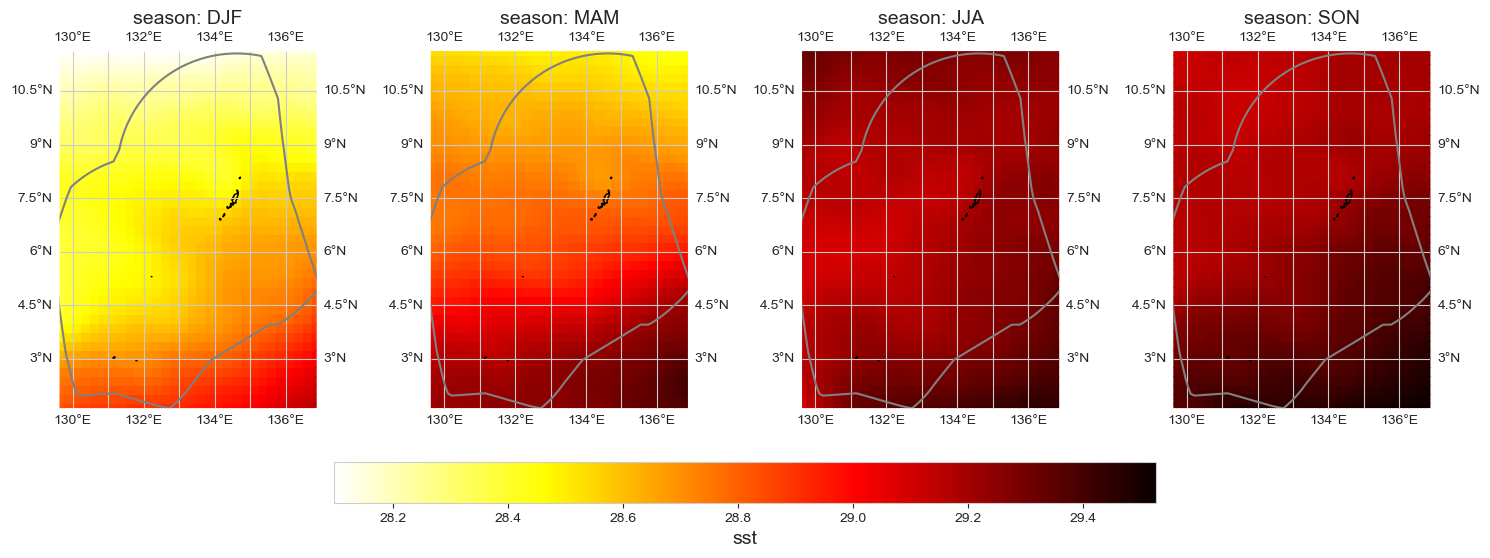
Seasonal average#
data_month = data_xr.groupby('time.season').mean().sel(season = ['DJF', 'MAM', 'JJA', 'SON'])
im = plot_map_subplots(data_month, dataset_id, shp_eez = shp_eez, cmap = 'hot_r',
vmin = np.nanpercentile(data_month.min(dim = 'season')[dataset_id], 1),
vmax = np.nanpercentile(data_month.max(dim = 'season')[dataset_id], 99.99),
figsize = (15,11), sub_plot = [1, 4], cbar_pad = 0.05,
cbar = 1)
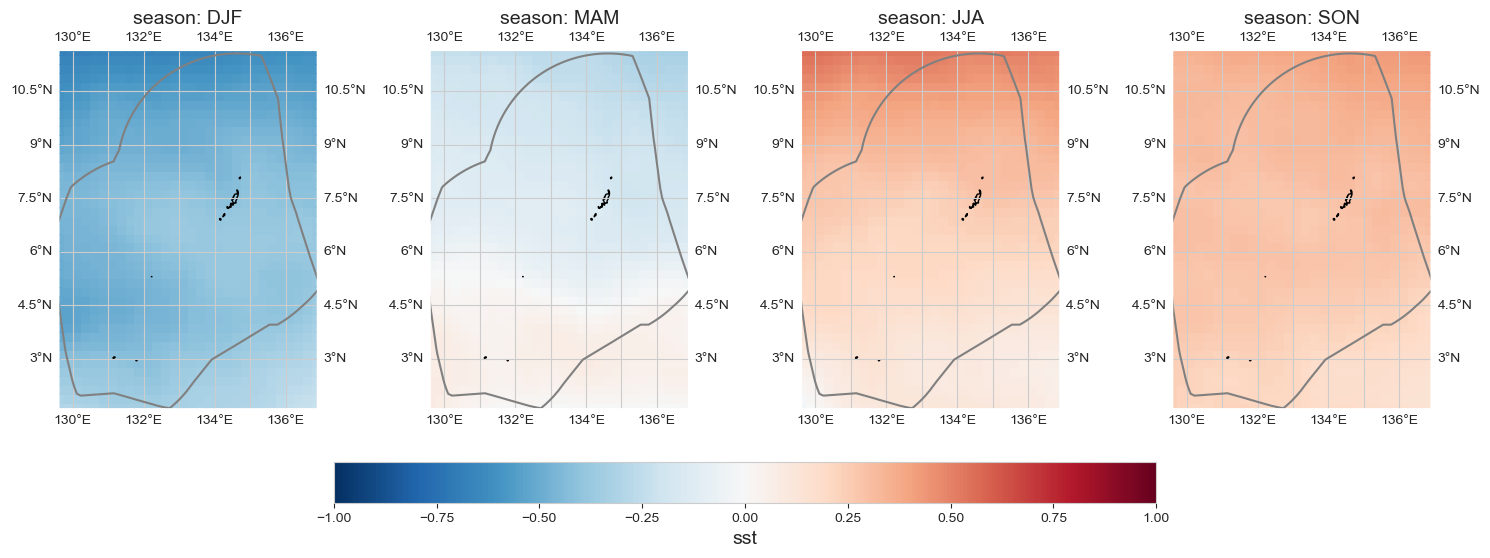
Seasonal anomaly#
data_month = data_xr.groupby('time.season').mean().sel(season = ['DJF', 'MAM', 'JJA', 'SON']) - data_xr.mean(dim='time')
im = plot_map_subplots(data_month, dataset_id, shp_eez = shp_eez,
cmap = 'RdBu_r', vmin=-1, vmax=1,
figsize = (15,11), sub_plot = [1, 4], cbar_pad = 0.05,
cbar = 1)
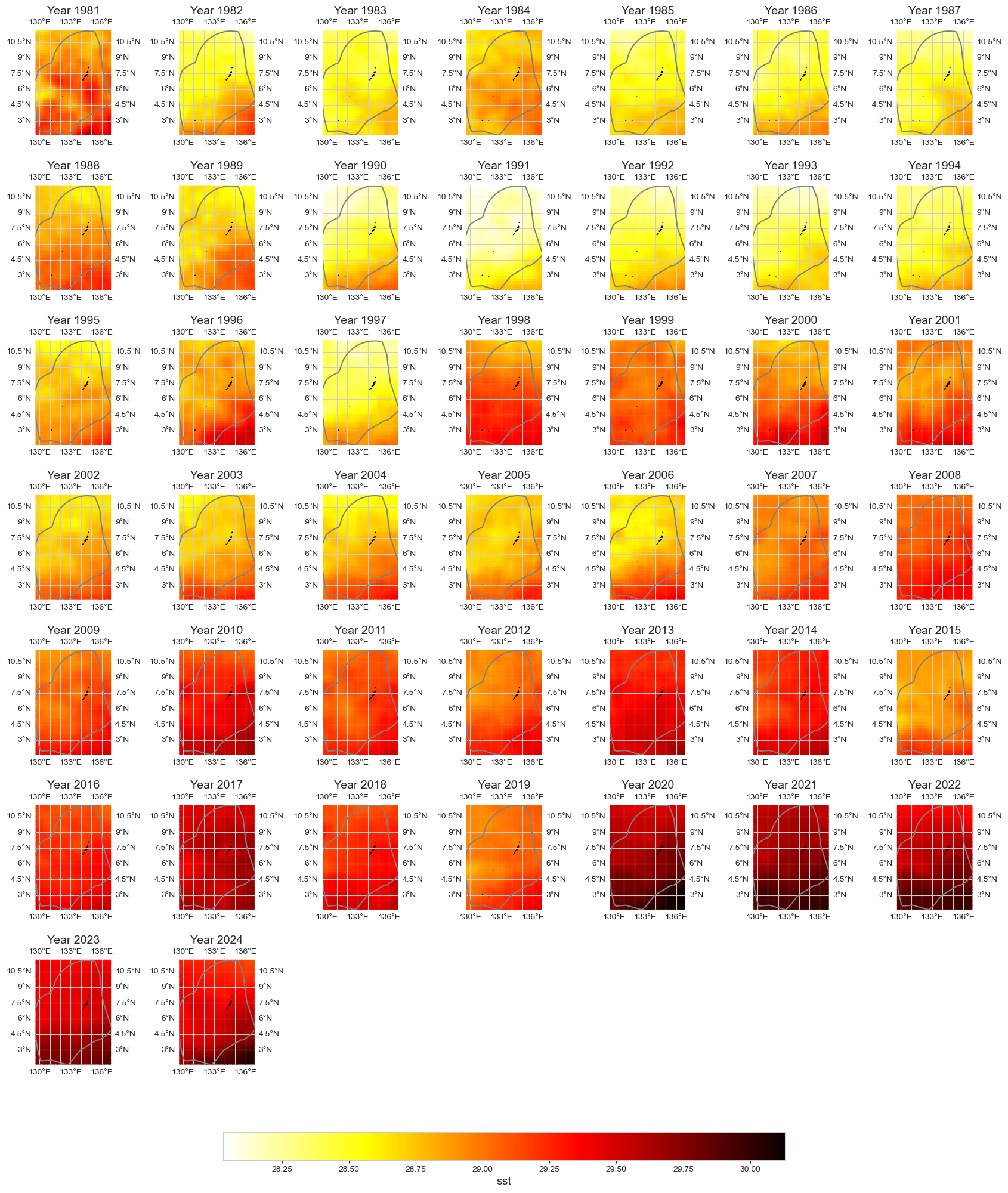
Annual average#
data_y = data.resample(time='1YE').mean()
im = plot_map_subplots(data_y, dataset_id, shp_eez = shp_eez, cmap = 'hot_r',
vmin = np.nanpercentile(data_y.min(dim = 'time')[dataset_id], 1),
vmax = np.nanpercentile(data_y.max(dim = 'time')[dataset_id], 99),
cbar = 1, cbar_pad = 0.05)
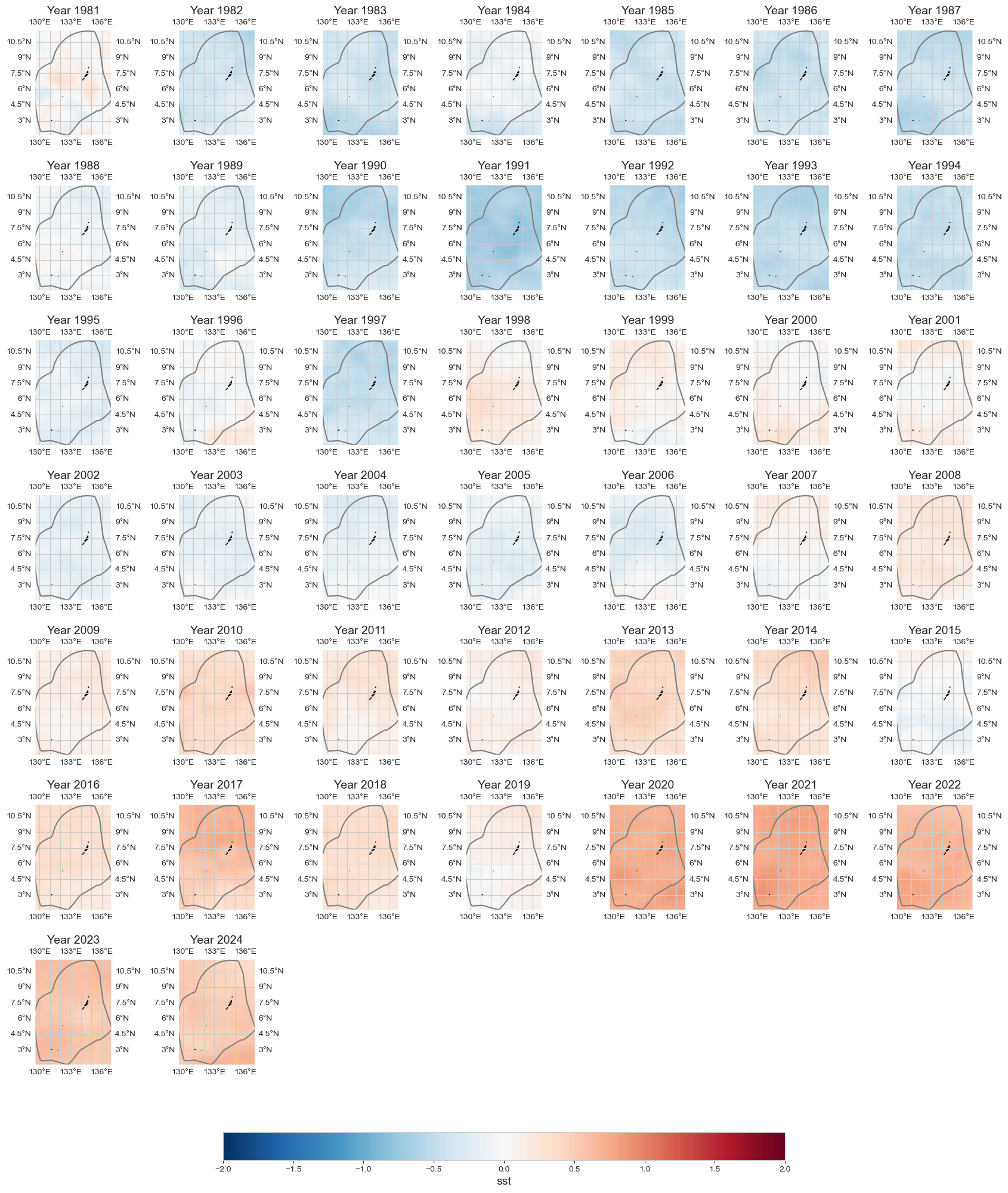
Annual anomaly#
data_an = data_y - data.mean(dim='time')
fig = plot_map_subplots(data_an, dataset_id, shp_eez = shp_eez, cmap='RdBu_r',
vmin=-2, vmax=2, cbar = 1, cbar_pad = 0.05)

Average Area#
data_mean = data.mean(dim = ['lon', 'lat']).to_dataframe()
datag = data_mean.groupby(data_mean.index.year).max()
datag.index = pd.to_datetime(datag.index, format = '%Y')
datag['sst_ref'] = datag['sst'] - datag.loc['1961':'1990'].sst.mean()
dict_plot = [{'data' : data.mean(dim = ['lon', 'lat']).to_dataframe(),
'var' : dataset_id, 'ax' : 1, 'label' : 'SST [ºC]'},]
fig = plot_timeseries_interactive(dict_plot, trendline=True, scatter_dict = None, figsize = (25, 12), label_yaxes = 'SST [ºC]');
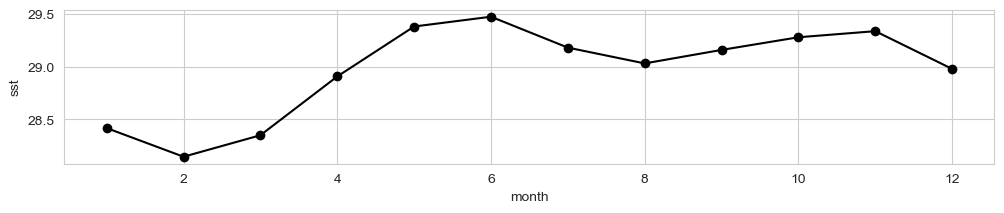
Seasonal average#
fig, ax = plt.subplots(figsize=(12,2))
data_xr.mean(dim = ['lon', 'lat']).groupby('time.month').mean()[dataset_id].plot(ax = ax, marker = 'o', color = 'k')
[<matplotlib.lines.Line2D at 0x188ed2900>]
nevents = 10
data_mean = data.mean(dim = ['lon', 'lat']).to_dataframe()
data_mean = data_mean.resample('YE').mean()
top_10 = data_mean.sort_values(by='sst', ascending=False).head(nevents)
fig, ax, trend = plot_bar_probs(x=data_mean.index.year, y=data_mean.sst, trendline=True,
y_label='SST [°C]', figsize=[15, 4], return_trend=True)
ax.set_ylim(data_mean.sst.min()-.2, data_mean.sst.max()+.2)
im = ax.scatter(top_10.index.year, top_10.sst,
c=top_10.sst.values, s=100, ec = 'pink', cmap='rainbow', label='Top 10 warmest years')
plt.colorbar(im, pad = 0).set_label('Absolute SST [°C]', fontsize=fontsize)
ax.set_title('Annual Mean SST', fontsize=15);
glue("average_timeseries", fig, display=False)
plt.savefig(op.join(path_figs, 'F12_SST_trends_Annomalies.png'), dpi=300, bbox_inches='tight')

Given point#
loc = [7.37, 134.7]
dict_plot = [{'data' : data.sel(lon=loc[1], lat=loc[0], method='nearest').to_dataframe(),
'var' : dataset_id, 'ax' : 1, 'label' : 'SST [ºC]'},]
fig, ax = plot_base_map(shp_eez = shp_eez, figsize = [10, 6])
ax.set_extent([lon_range[0], lon_range[1], lat_range[0], lat_range[1]], crs=ccrs.PlateCarree())
ax.plot(loc[1], loc[0], '*', markersize = 12, color = 'royalblue', transform=ccrs.PlateCarree(), label = 'Location Analysis')
ax.legend()
<matplotlib.legend.Legend at 0x188ebc560>
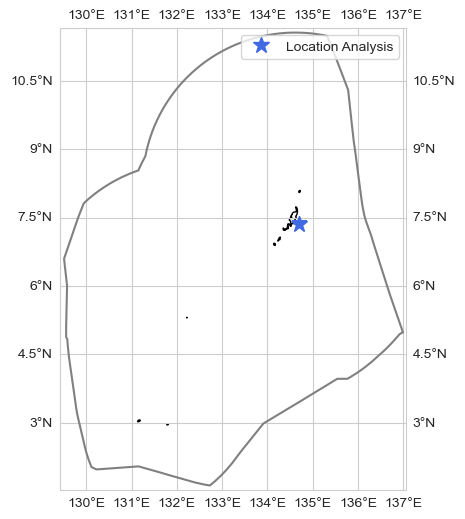
fig = plot_timeseries_interactive(dict_plot, trendline=True, scatter_dict = None, figsize = (25, 12), label_yaxes = 'SST [ºC]');
ONI index analysis#
p_data = 'https://psl.noaa.gov/data/correlation/oni.data'
df1 = download_oni_index(p_data)
lims = [-.5, .5]
plot_oni_index_th(df1, lims = lims)
Group by ONI category
data_xr = xr.load_dataset(op.join(path_data, 'sst_daily_1981_2024_palau.nc')).rename({'lat':'latitude', 'lon':'longitude'}).resample(time='1M').mean()
data_xr['time'] = data_xr.time.values.astype('datetime64[M]')
data_xr['ONI'] = (('time'), df1.iloc[np.intersect1d(data_xr.time, df1.index, return_indices=True)[2]].ONI.values)
data_xr['ONI_cat'] = (('time'), np.where(data_xr.ONI < lims[0], -1, np.where(data_xr.ONI > lims[1], 1, 0)))
data_oni = data_xr.groupby('ONI_cat').mean()
data_oni['labels'] = (('ONI_cat'),['La Niña', 'Neutral', 'El Niño'])
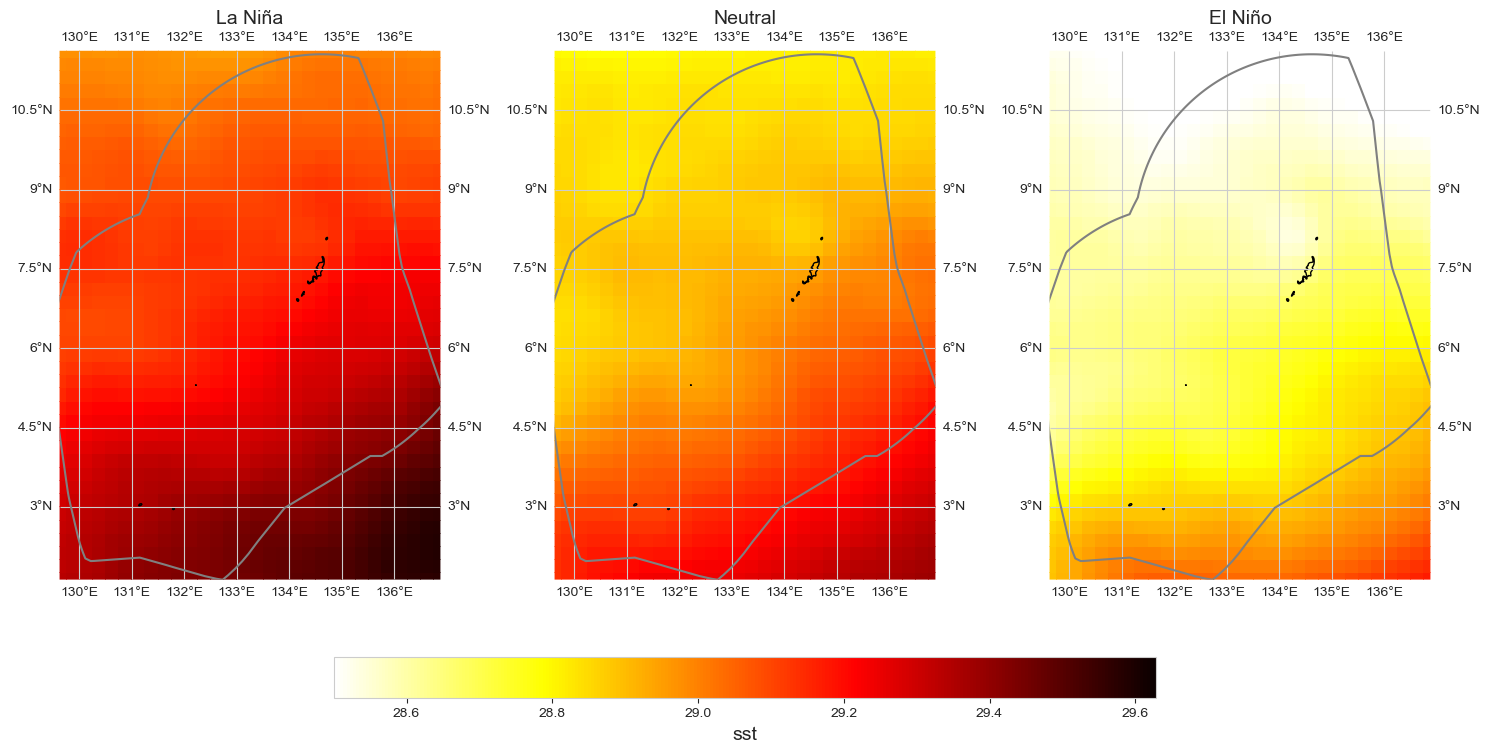
dataset_id = 'sst'
fig = plot_map_subplots(data_oni, dataset_id, shp_eez = shp_eez, cmap = 'hot_r',
vmin = np.percentile(data_xr.mean(dim = 'time')[dataset_id], 0) - .25,
vmax = np.percentile(data_xr.mean(dim = 'time')[dataset_id], 100) + .25,
sub_plot= [1, 3], figsize = (15, 8), cbar = True, cbar_pad = .1)
plt.savefig(op.join(path_figs, 'F14_SST_ENSO.png'), dpi=300, bbox_inches='tight')
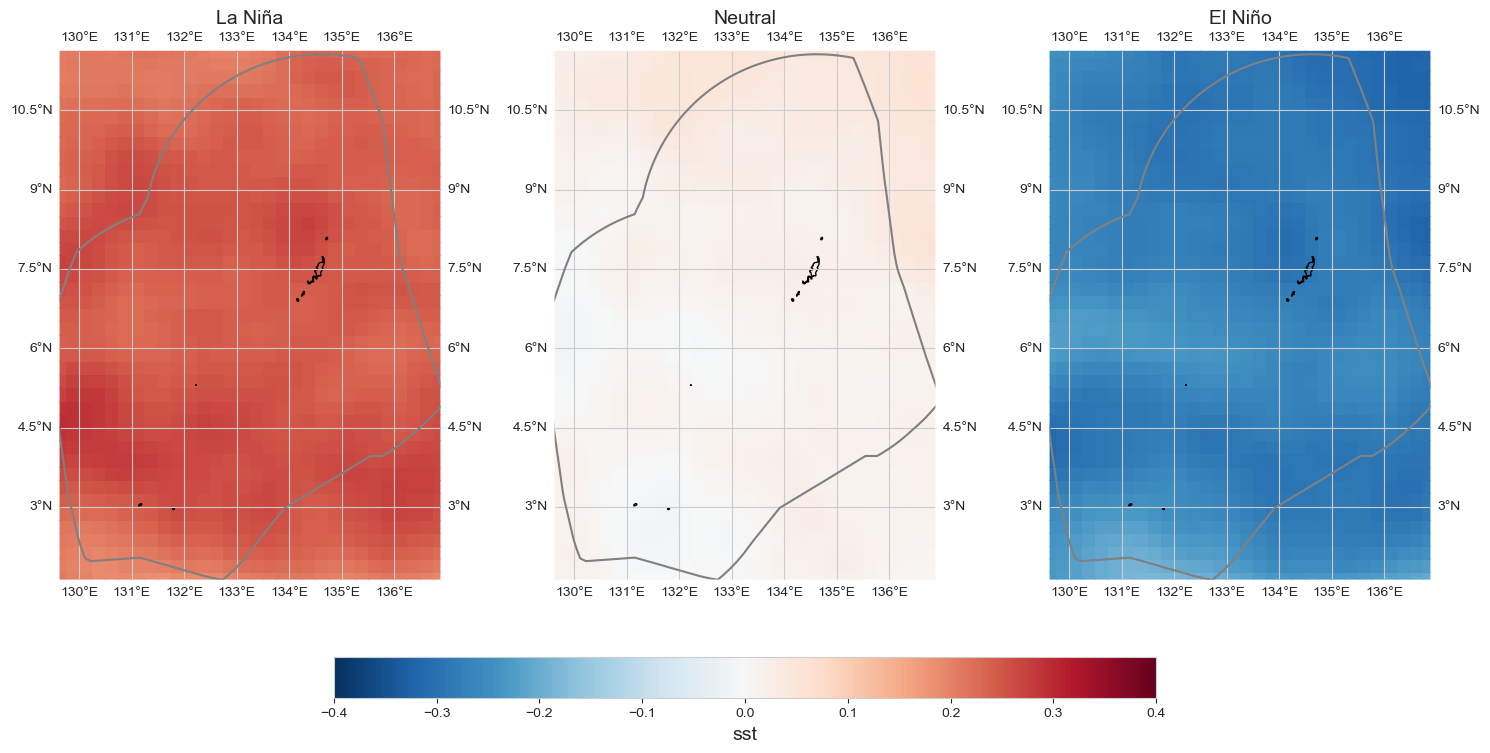
data_an = data_oni - data_xr.mean(dim='time')
data_an['labels'] = (('ONI_cat'),['La Niña', 'Neutral', 'El Niño'])
fig = plot_map_subplots(data_an, dataset_id, shp_eez = shp_eez, cmap='RdBu_r', vmin=-.4, vmax=.4,
sub_plot= [1, 3], figsize = (15, 8), cbar = True, cbar_pad = 0.1)
Table#
from ind_setup.tables import style_matrix, table_ocean
style_matrix(table_ocean(data, trend, data_oni, 'sst'))
| Metric | Value |
|---|---|
| Daily Average | 28.973 |
| Daily Maximum 21/09/1998 | 30.812 |
| Daily Minimum 03/03/1992 | 26.710 |
| Maximum Annual Average | 29.736 |
| Minimum Annual Average | 28.300 |
| Rate of change [°C/year] | 0.024 |
| Change between 1981 and 2024 [°C] | 1.032 |
| Average La Niña sst | 29.213 |
| Average El Niño sst | 28.697 |
| Average Neutral sst | 28.988 |
