Estimated Phytoplankton Size#
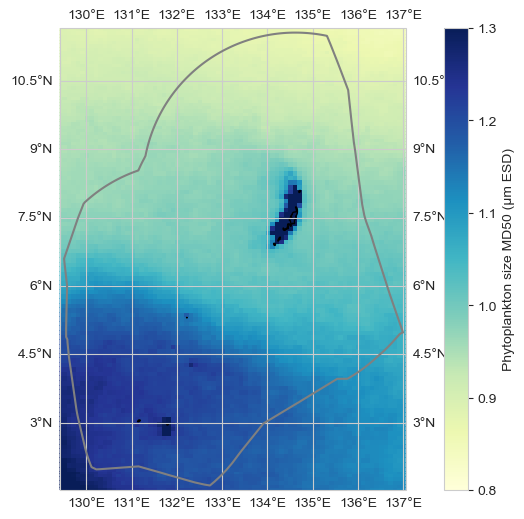
Figure. Estimated Median Phytoplankton Size from satellite. The map (top) shows the change in the estimated median phytoplankton size (μm equivalent spherical diameter, ESD) in the vicinity of Palau over the period 1998-2023 derived from satellite remotely sensed sea surface temperature and ocean color data. The grey line is the Palau EEZ. The line plot (bottom) shows the change in the estimated median phytoplankton size (μm ESD) averaged over the area within the top plot. The black line represents the trend, which is not statistically significant (p > 0.05).
Show code cell source
import warnings
warnings.filterwarnings("ignore")
import os
import os.path as op
import sys
import pandas as pd
import numpy as np
import xarray as xr
import geopandas as gpd
import cartopy.crs as ccrs
import matplotlib.pyplot as plt
from myst_nb import glue
sys.path.append("../../../../indicators_setup")
from ind_setup.plotting_int import plot_timeseries_interactive, plot_oni_index_th
from ind_setup.plotting import plot_base_map, plot_map_subplots, add_oni_cat, plot_bar_probs, fontsize
sys.path.append("../../../functions")
from data_downloaders import download_ERDDAP_data, download_oni_index
from ocean import process_trend_with_nan
Setup#
Define area of interest
#Area of interest
lon_range = [129.4088, 137.0541]
lat_range = [1.5214, 11.6587]
EEZ shapefile
shp_f = op.join(os.getcwd(), '..', '..','..', 'data/Palau_EEZ/pw_eez_pol_april2022.shp')
shp_eez = gpd.read_file(shp_f)
Download Data#
DATASET: https://oceanwatch.pifsc.noaa.gov/erddap/info/md50_exp/index.html
update_data = False
path_data = "../../../data"
path_figs = "../../../matrix_cc/figures"
Show code cell source
base_url = 'https://oceanwatch.pifsc.noaa.gov/erddap/griddap/md50_exp.csv'
dataset_id = 'MD50'
if update_data:
date_ini = '1998-01-01T00:00:00Z'
date_end = '2024-12-01T00:00:00Z'
data = download_ERDDAP_data(base_url, dataset_id, date_ini, date_end, lon_range, lat_range)
data_xr = data.set_index(['latitude', 'longitude', 'time']).to_xarray()
data_xr['time'] = pd.to_datetime(data_xr.time)
data_xr = data_xr.coarsen(longitude=2, latitude=2, boundary = 'pad').mean()
data_xr.to_netcdf(op.join(path_data, f'griddap_{dataset_id}.nc'))
else:
data_xr = xr.open_dataset(op.join(path_data, f'griddap_{dataset_id}.nc'))
Analysis#
Plotting#
Average#
fig, ax = plot_base_map(shp_eez = shp_eez, figsize = [10, 6])
im = ax.pcolor(data_xr.longitude, data_xr.latitude, data_xr.mean(dim='time')[dataset_id], transform=ccrs.PlateCarree(),
cmap = 'YlGnBu', vmin = 0.8, vmax = 1.3)
ax.set_extent([lon_range[0], lon_range[1], lat_range[0], lat_range[1]], crs=ccrs.PlateCarree())
plt.colorbar(im, ax=ax, label='Phytoplankton size MD50 (µm ESD)')
glue("average_map", fig, display=False)
plt.savefig(op.join(path_figs, 'F16_phytoplankton_mean_map.png'), dpi=300, bbox_inches='tight')

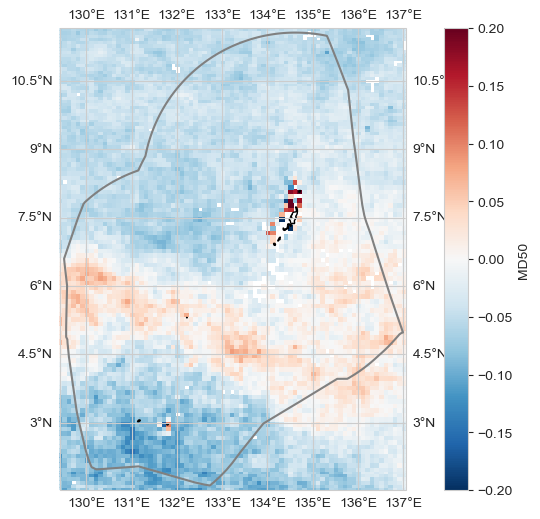
Change#
trend_m, _, _, _, _ = process_trend_with_nan(data_xr[dataset_id])
fig, ax = plot_base_map(shp_eez = shp_eez, figsize = [10, 6])
im = ax.pcolor(data_xr.longitude, data_xr.latitude,
trend_m,
transform=ccrs.PlateCarree(),
cmap = 'RdBu_r',
vmin = -.2,
vmax = .2,
)
ax.set_extent([lon_range[0], lon_range[1], lat_range[0], lat_range[1]], crs=ccrs.PlateCarree())
plt.colorbar(im, ax=ax, label= dataset_id)
plt.savefig(op.join(path_figs, 'F16_phytoplankton_mean_map_trend.png'), dpi=300, bbox_inches='tight')
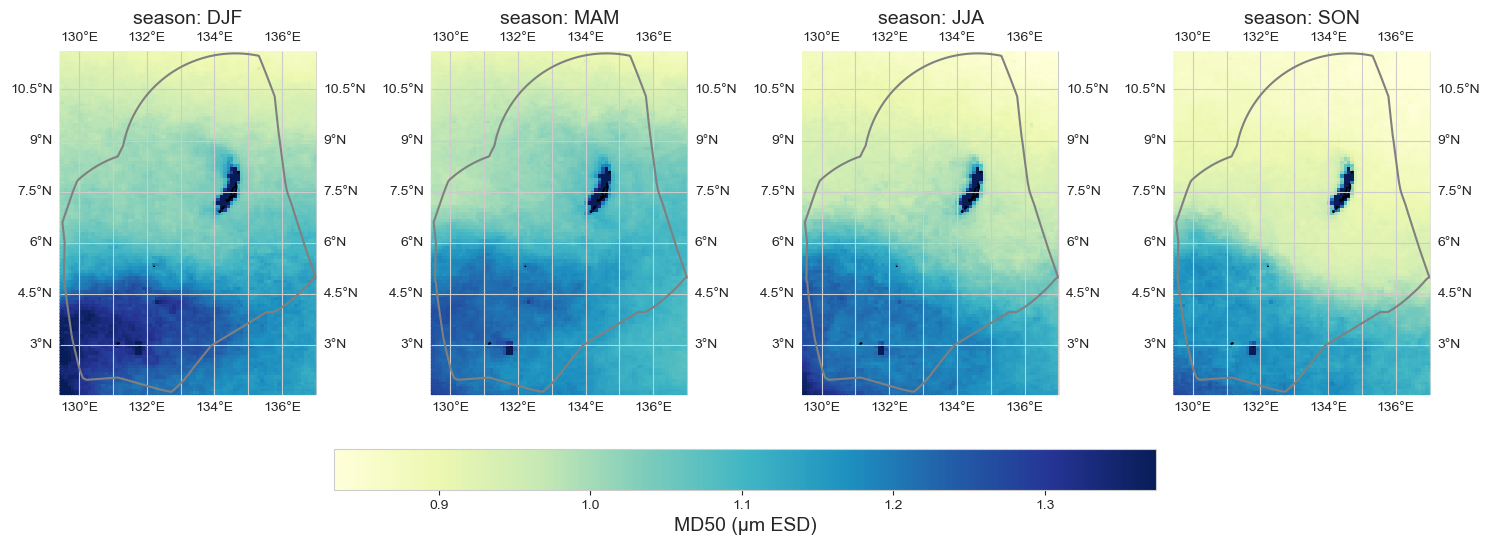
Seasonal average#
data_month = data_xr.groupby('time.season').mean().sel(season = ['DJF', 'MAM', 'JJA', 'SON'])
im = plot_map_subplots(data_month, dataset_id, shp_eez = shp_eez, cmap = 'YlGnBu',
vmin = np.nanpercentile(data_month.min(dim = 'season')[dataset_id], 1),
vmax = np.nanpercentile(data_month.max(dim = 'season')[dataset_id], 99),
figsize = (15,11), sub_plot = [1, 4], cbar_pad = 0.05, cbar = 1)
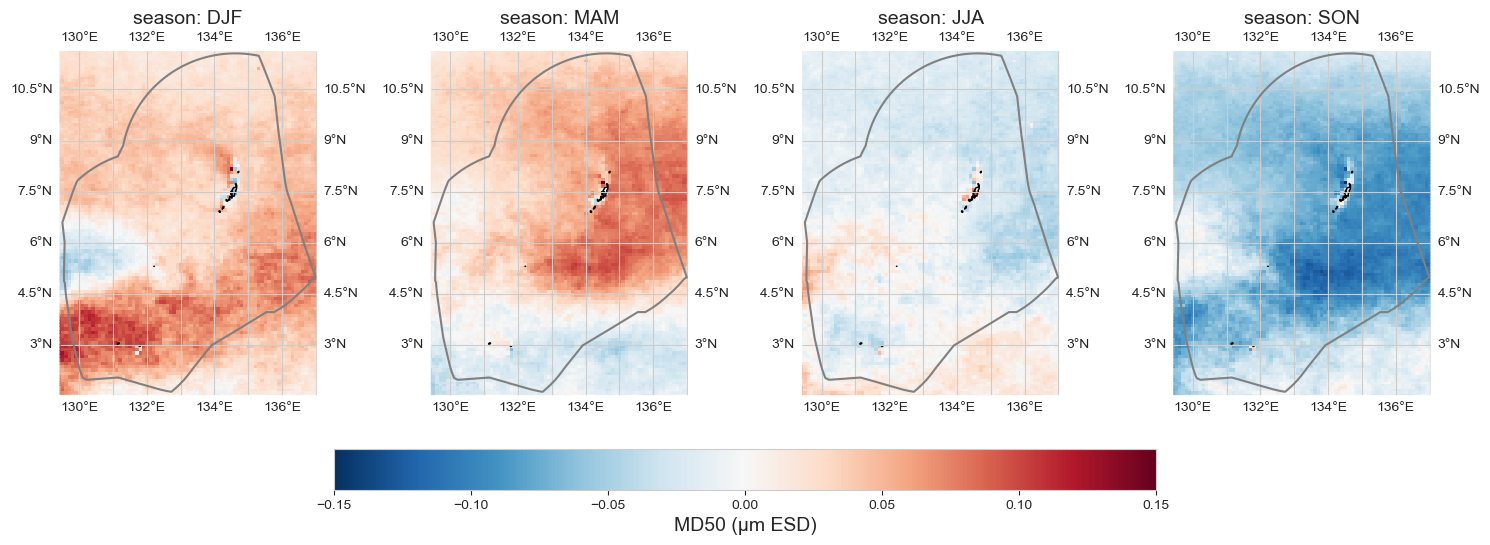
Seasonal anomaly#
data_month = data_xr.groupby('time.season').mean().sel(season = ['DJF', 'MAM', 'JJA', 'SON']) - data_xr.mean(dim='time')
im = plot_map_subplots(data_month, dataset_id, shp_eez = shp_eez,
cmap = 'RdBu_r', vmin=-.15, vmax=.15,
figsize = (15,11), sub_plot = [1, 4], cbar_pad = 0.05,
cbar = 1)
Annual average#
data_y = data_xr.resample(time='1YE').mean()
fig = plot_map_subplots(data_y, 'MD50', shp_eez = shp_eez, cmap = 'YlGnBu', vmin = 0.4, vmax = 1.6, cbar = 1)
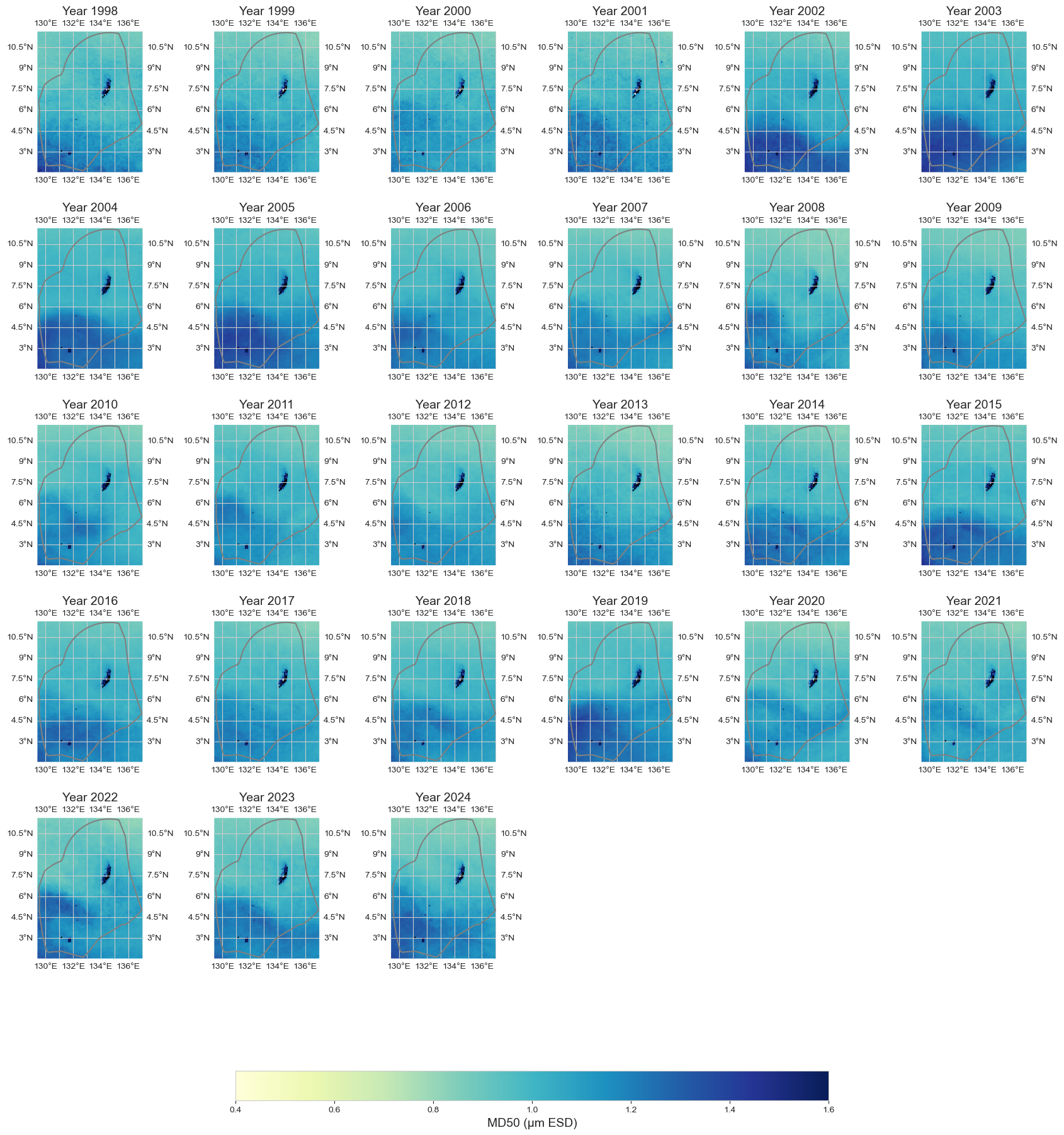
Annual anomaly#
data_an = data_y - data_xr.mean(dim='time')
fig = plot_map_subplots(data_an, dataset_id, shp_eez = shp_eez, cmap='RdBu_r', vmin=-.3, vmax=.3, cbar = 1)
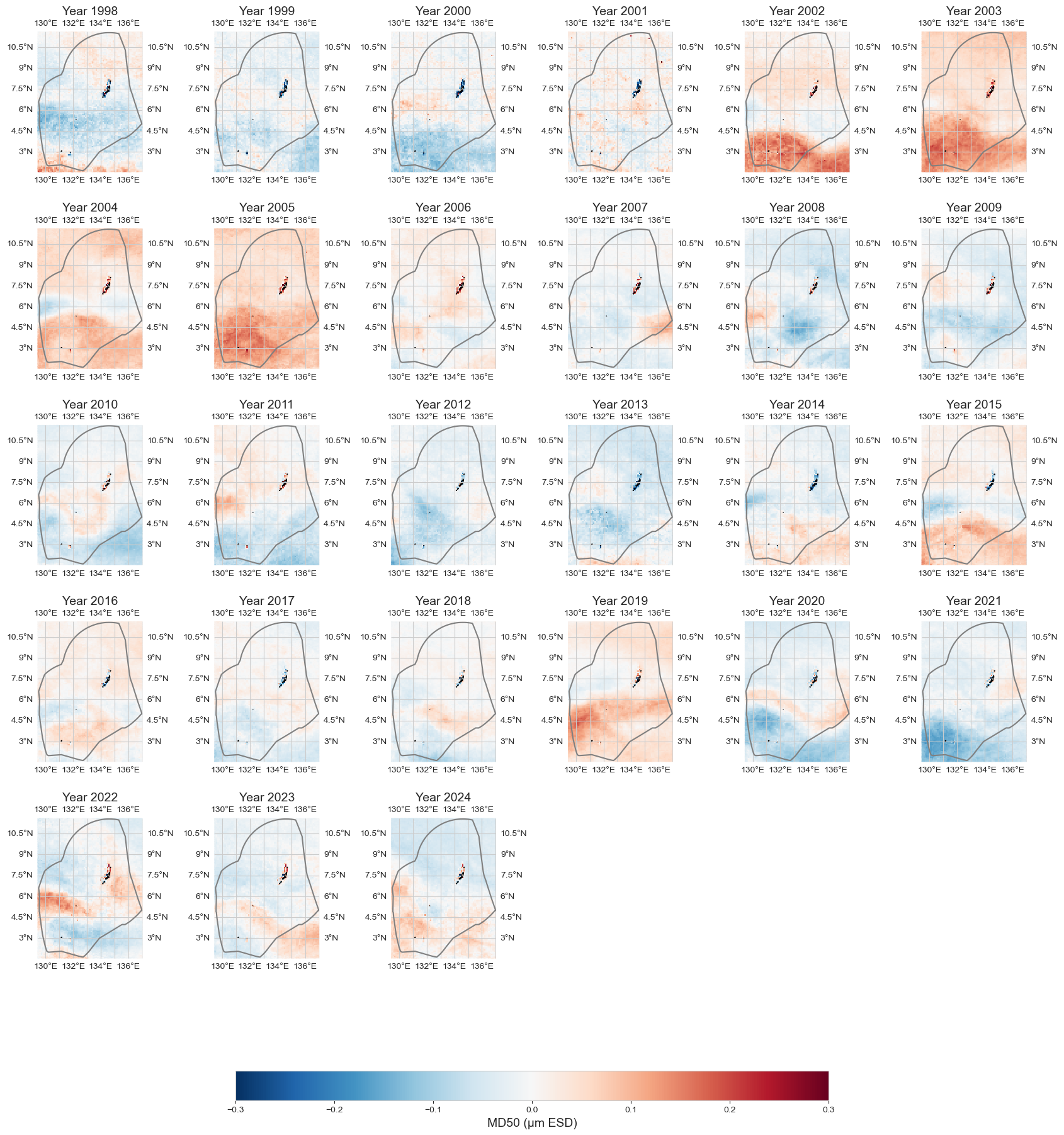
Average over area#
dict_plot = [{'data' : data_xr.mean(dim = ['longitude', 'latitude']).to_dataframe(),
'var' : dataset_id, 'ax' : 1, 'label' : 'Median Phytoplankton Size - MEAN AREA'},]
fig, trend = plot_timeseries_interactive(dict_plot, trendline=True, return_trend=True, scatter_dict = None, figsize = (25, 12),
label_yaxes = 'Phytoplankton MD50 (µm ESD)');
fig.write_html(op.join(path_figs, 'F16_phytoplankton_mean_trend.html'), include_plotlyjs="cdn")
Seasonal variability#
fig, ax = plt.subplots(figsize=(12,2))
data_xr.mean(dim = ['longitude', 'latitude']).groupby('time.month').mean()[dataset_id].plot(ax = ax, marker = 'o', color = 'k')
[<matplotlib.lines.Line2D at 0x19023b1d0>]
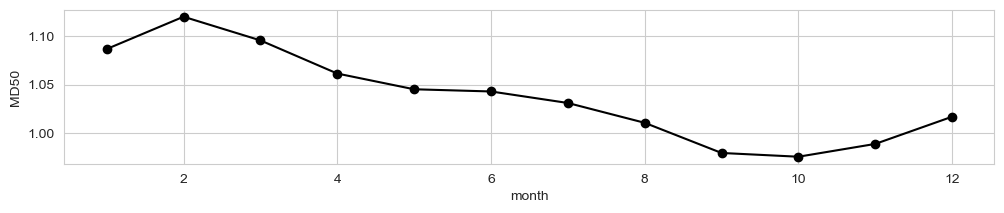
d_p = data_xr.mean(dim = ['longitude', 'latitude']).to_dataframe()
Timeseries at a given point#
loc = [7.37, 134.7]
dict_plot = [{'data' : data_xr.sel(longitude=loc[1], latitude=loc[0], method='nearest').to_dataframe(),
'var' : dataset_id, 'ax' : 1, 'label' : f'Median Phytoplankton Size at [{loc[0]}, {loc[1]}]'},]
fig, ax = plot_base_map(shp_eez = shp_eez, figsize = [10, 6])
ax.set_extent([lon_range[0], lon_range[1], lat_range[0], lat_range[1]], crs=ccrs.PlateCarree())
ax.plot(loc[1], loc[0], '*', markersize = 12, color = 'royalblue', transform=ccrs.PlateCarree(), label = 'Location Analysis')
ax.legend()
<matplotlib.legend.Legend at 0x1903a95b0>
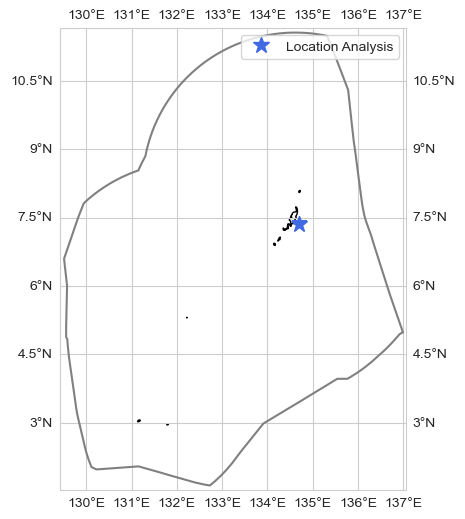
fig = plot_timeseries_interactive(dict_plot, trendline=True, scatter_dict = None, figsize = (25, 12),
label_yaxes = 'Phytoplankton MD50 (µm ESD)');
ONI index analysis#
if update_data:
p_data = 'https://psl.noaa.gov/data/correlation/oni.data'
df1 = download_oni_index(p_data)
df1.to_pickle(op.join(path_data, 'oni_index.pkl'))
else:
df1 = pd.read_pickle(op.join(path_data, 'oni_index.pkl'))
lims = [-.5, .5]
plot_oni_index_th(df1, lims = lims)
Group by ONI category
df1 = add_oni_cat(df1, lims = lims)
df1['ONI'] = df1['oni_cat']
data_xr['ONI'] = (('time'), df1.iloc[np.intersect1d(data_xr.time, df1.index, return_indices=True)[2]].ONI.values)
data_xr['ONI_cat'] = (('time'), np.where(data_xr.ONI < lims[0], -1, np.where(data_xr.ONI > lims[1], 1, 0)))
data_oni = data_xr.groupby('ONI_cat').mean()
Average#
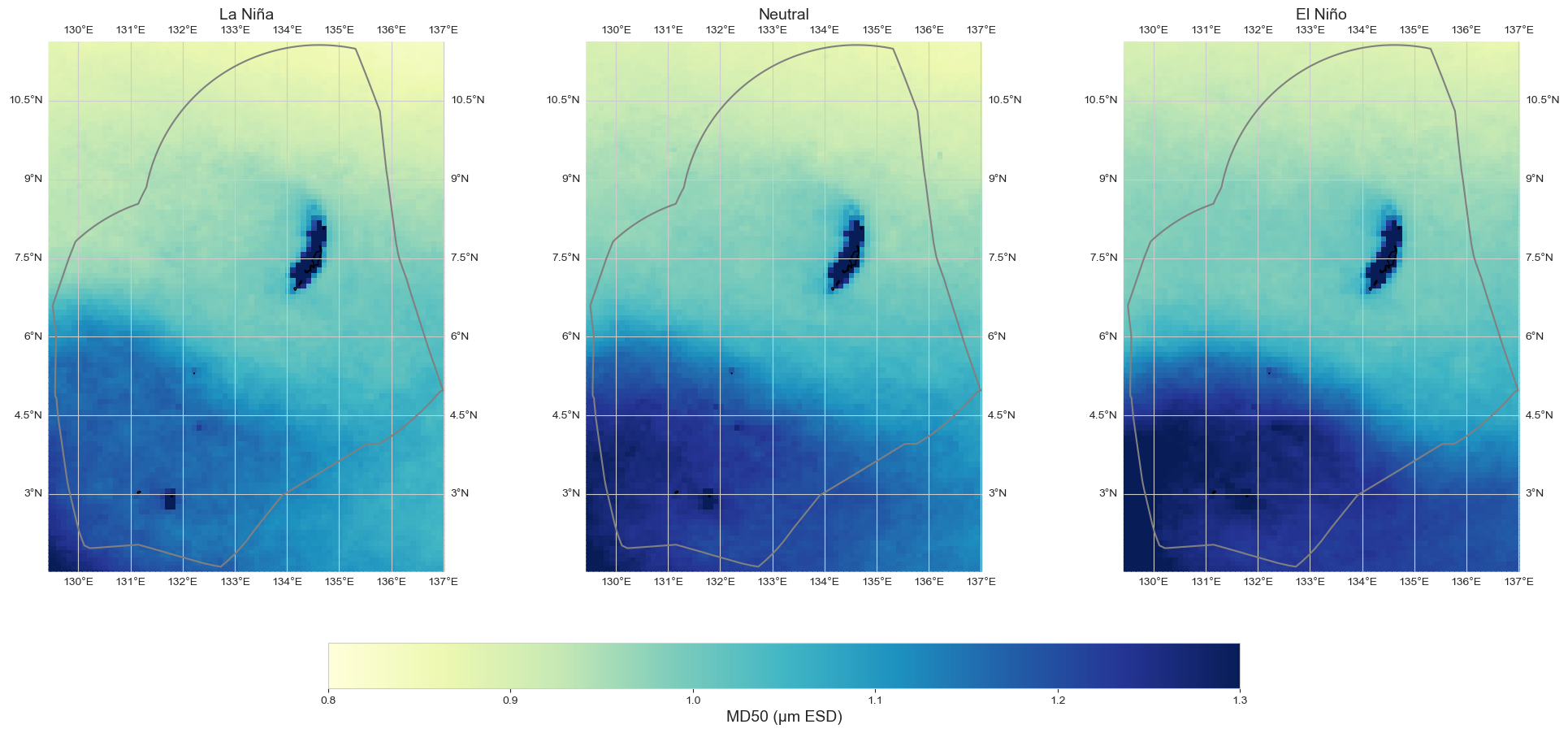
fig = plot_map_subplots(data_oni, dataset_id, shp_eez = shp_eez, cmap = 'YlGnBu', vmin = 0.8, vmax = 1.3,
sub_plot= [1, 3], figsize = (20, 9), cbar = True, cbar_pad = 0.1,
titles = ['La Niña', 'Neutral', 'El Niño'],)
plt.savefig(op.join(path_figs, 'F16_phytoplankton_ENSO.png'), dpi=300, bbox_inches='tight')
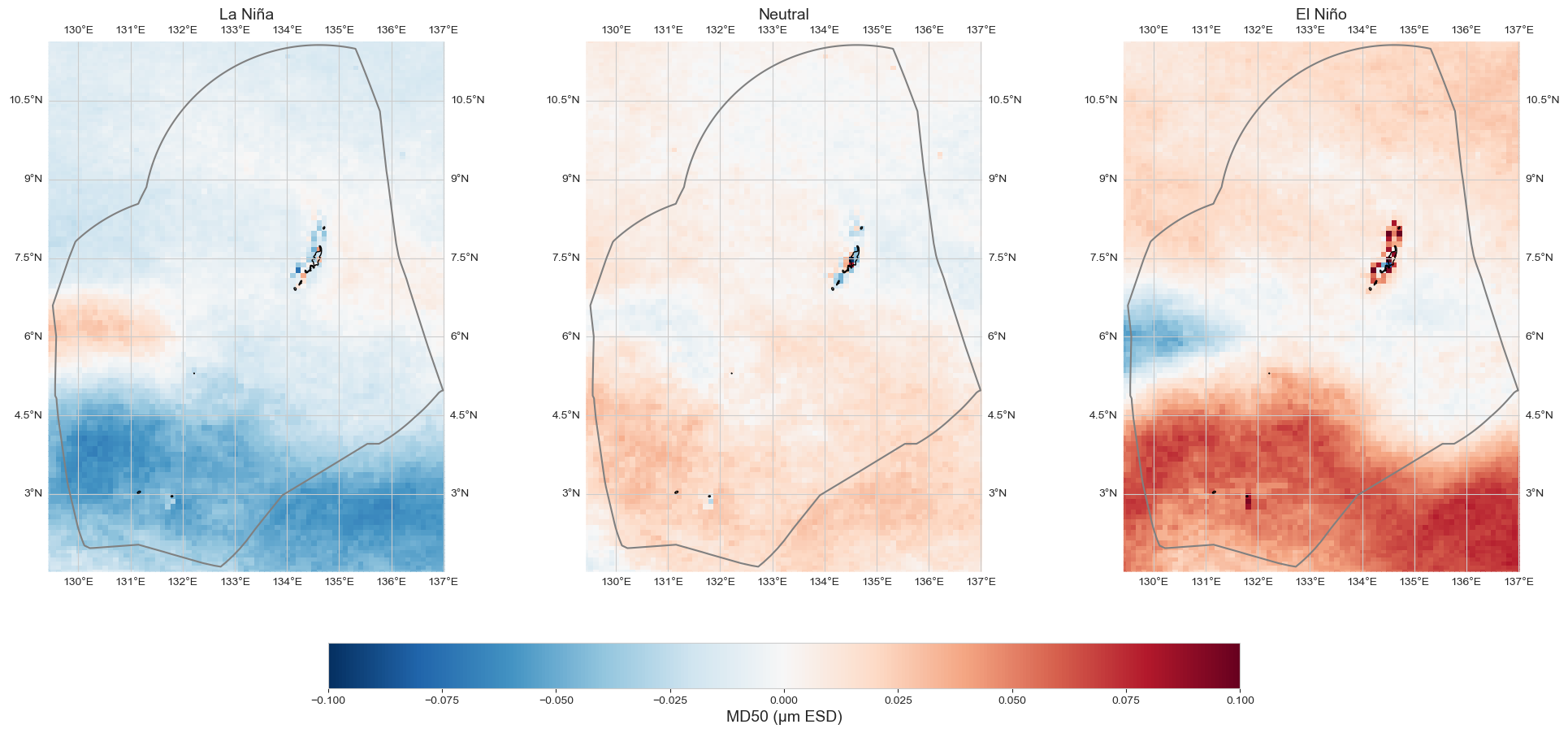
Anomaly#
data_an = data_oni - data_xr.mean(dim='time')
fig = plot_map_subplots(data_an, dataset_id, shp_eez = shp_eez, cmap='RdBu_r', vmin=-.1, vmax=.1,
sub_plot= [1, 3], figsize = (20, 9), cbar = True, cbar_pad = 0.1,
titles = ['La Niña', 'Neutral', 'El Niño'],)
Table#
from ind_setup.tables import style_matrix, table_ocean
style_matrix(table_ocean(data_xr, trend[0], data_oni, dataset_id))
| Metric | Value |
|---|---|
| Monthly Average | 1.038 |
| Monthly Maximum 01/02/2005 | 1.266 |
| Monthly Minimum 01/09/1998 | 0.821 |
| Maximum Annual Average | 1.113 |
| Minimum Annual Average | 0.998 |
| Rate of change [µm ESD/year] | -0.001 |
| Change between 1998 and 2024 [µm ESD] | -0.026 |
| Average La Niña MD50 | 1.020 |
| Average El Niño MD50 | 1.061 |
| Average Neutral MD50 | 1.046 |
